My research lies at the intersection of biological imaging, fluorescence microscopy, and computational image analysis. I develop methods to visualize and quantify sub-cellular processes with improved spatial and temporal resolution, tackling challenges in big data analysis, 4D visualization, and automated quantification.
Currently, I focus on two main directions: studying the cell division mechanisms of cyanobacteria through deep learning-based image analysis, and advancing quantitative nanoscale imaging of molecular orientation through polarized super-resolution microscopy.
Cell division of cyanobacteria
Deep learning-based algorithms for cell segmentation and tracking
About: I develop automatic tracking methods for single cells, filaments, and cell populations of cyanobacteria, combining advanced microscopy with deep learning to study cell division mechanisms and growth dynamics.
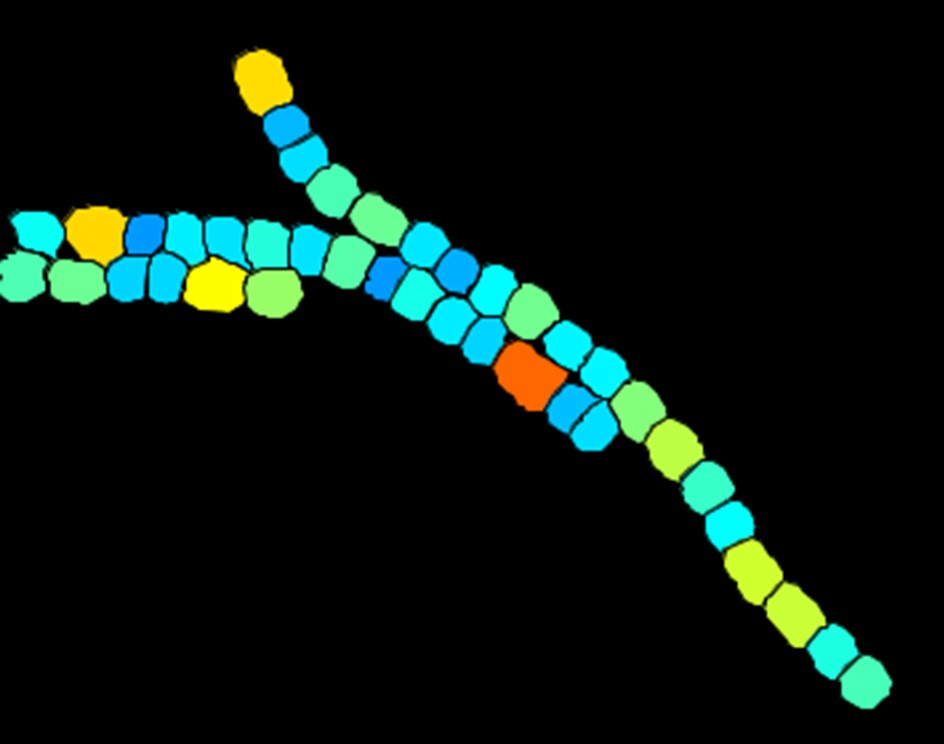 Cell segmentation
Cell segmentation
Quantitative nanoscale imaging of molecular orientation
Polarized super-resolution microscopy for imaging orientational order in biological structures
About: Long-standing collaboration with Sophie Brasselet’s team at Institut Fresnel, France. We develop polarization-based super-resolution methods to measure the nanoscale organization and 3D orientation of biological filaments such as actin networks.
Latest: Our new method 4polar3D, published in Nature Communications (2026), enables 3D orientation measurements of single molecules in dense actin structures using ratiometric polarization splitting — with minimal optical complexity and fast data processing. Read the paper →
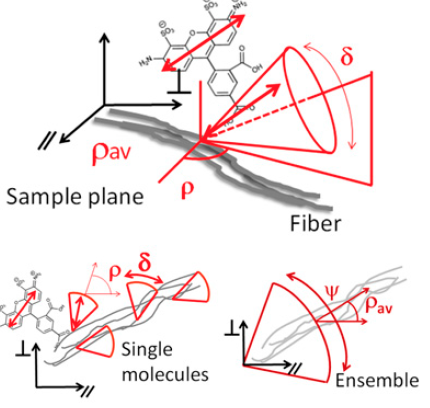 Ultrastructure imaging
Ultrastructure imaging
3D+time live cell imaging
NAVISCOPE: navigation and visualization of large-scale live cell imaging data
About: INRIA IPL project combining machine learning-driven detection of regions of interest with automatic quantification of molecular interactions and cell processes, enabling efficient navigation of large 3D+time datasets.
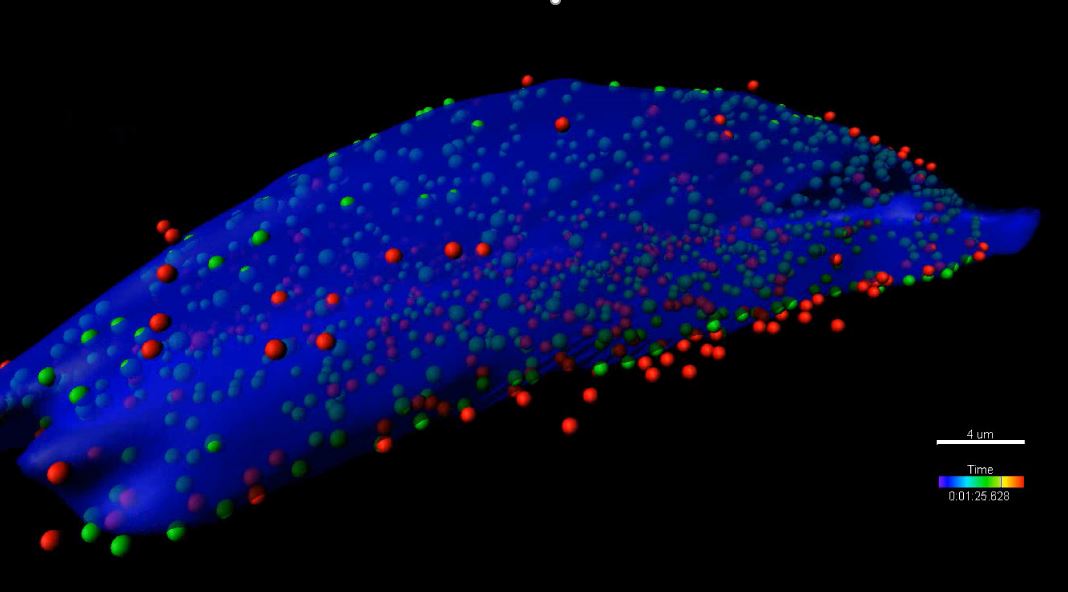 3D spots and cell segmentation
3D spots and cell segmentation
Classification of endocytic entry mechanisms
About: Analysis of clathrin-mediated and glyco(sphingo)lipid/lectin (GL-Lect)-mediated endocytosis using Lattice Light-Sheet Microscopy (LLSM) and 3D single particle tracking.
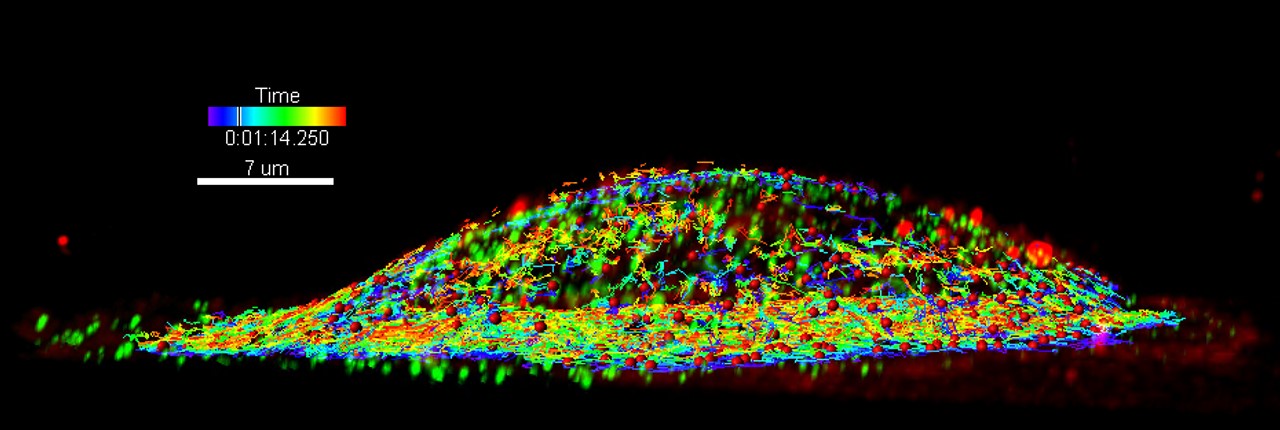 LLSM and 3D single particle tracking
LLSM and 3D single particle tracking
Image preprocessing and data management
BioImageIT: open-source framework for image data management and analysis
About: Project of the Serpico-STED Team within the National Research Infrastructure France BioImaging, providing an integrative platform for image data management and analysis across the 18 imaging facilities of the infrastructure.
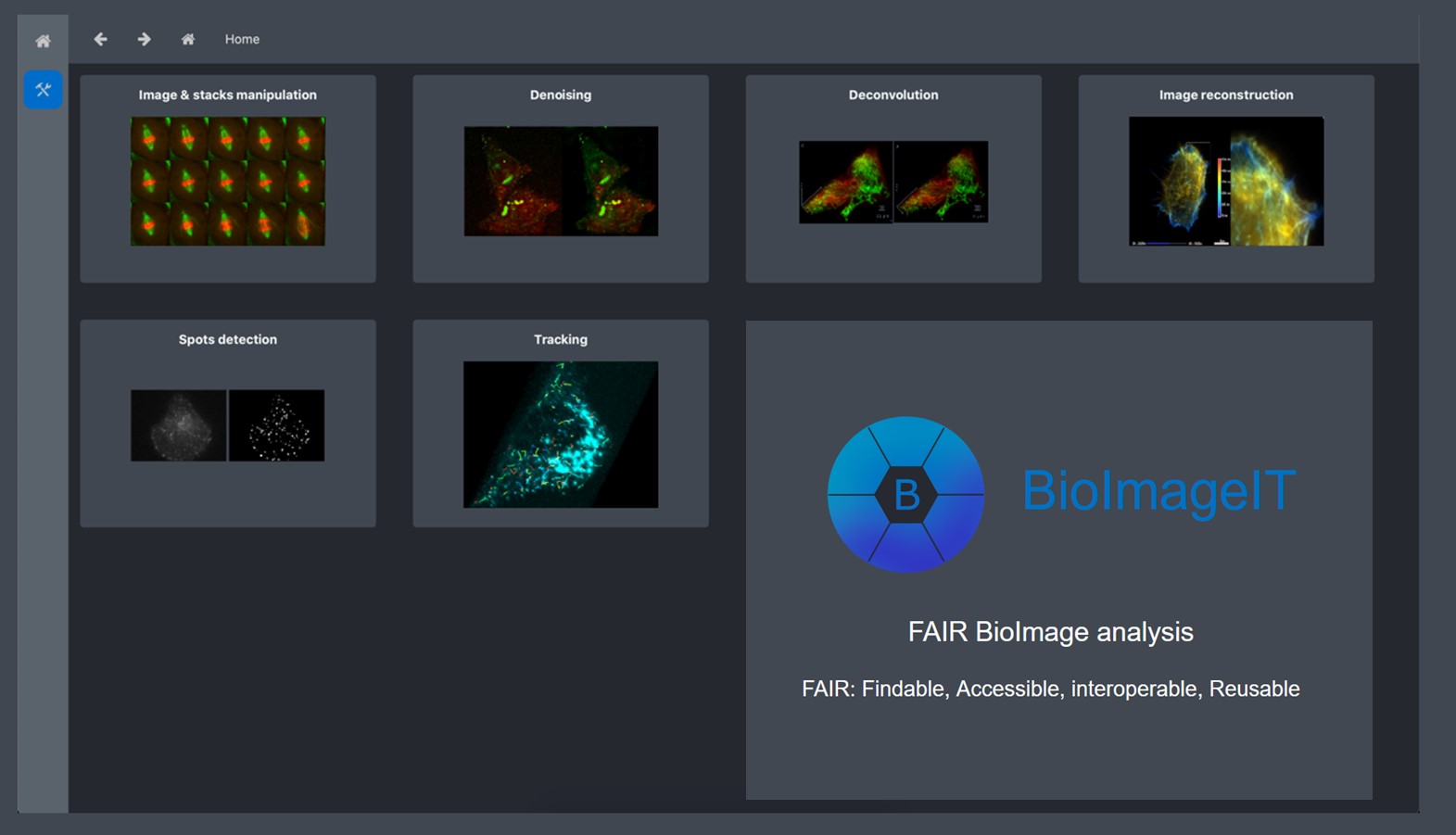 BioImageIT
BioImageIT
